2-Aminopurine can substitute for dA in an oligonucleotide. It is a naturally fluorescent base that is sensitive to the local environment making it a useful probe for monitoring the structure and dynamics of DNA hairpins and for detecting the base stacking state of a duplex. 2-Aminopurine can be destabilizing and slightly lower the Tm.
| Scientific Details |
| MW: |
313.2 |
| Mod Code: |
/52AmPr/ |
| Extinction Coefficient: |
1000 |
| Absorbance Max: |
303 nm |
| Emission Max: |
371 |
Extinction Coefficient
(At Absorbance max): |
3600 |
| Scientific Details |
| MW: |
313.2 |
| Mod Code: |
/i2AmPr/ |
| Extinction Coefficient: |
1000 |
| Absorbance Max: |
303 nm |
| Emission Max: |
371 |
Extinction Coefficient
(At Absorbance max): |
3600 |
| Scientific Details |
| MW: |
313.2 |
| Mod Code: |
/32AmPu/ |
| Extinction Coefficient: |
1000 |
| Absorbance Max: |
303 nm |
| Emission Max: |
371 |
Extinction Coefficient
(At Absorbance max): |
3600 |
Structure
Mix of 20 Trimers: (AAA+AAC+ACT+ATC+ATG+CAG+CAT+CCG+CGT+CTG+GAA+GAC+GCT+GGT+GTT+TAC+TCT+TGC+TGG+TTC) It is common practice to insert random N-base domains into oligonucleotides to perform mutagenesis or to evolve novel proteins (random sequence libraries). Unfortunately, inserting serial N bases gives rise to all 64 possible codons and thus does not produce an equal representation of the 20 amino acids (AAs). It will also insert unwanted stop codons. A set of trimer phosphoramidites have been developed which comprise a single codon for each of the 20 AAs. These are available as a 20 Trimer Mix for creating better N-domains into oligonucleotides intended to encode proteins. Each trimer addition inserts 3 bases into a DNA sequence. It is also possible to obtain custom mixes with more limited AA content. Note that the use of trimer mixes will not be useful to create N-base domains for aptamer evolution libraries.
This modified base can form three hydrogen bonds when base-paired with dT and can increase the Tm of short oligos by as much as 1-2ЁЦC per insertion. This effect, however, is complex and is dependent on sequence context.
| Scientific Details |
| MW: |
15 |
| Mod Code: |
/5AmdA/ |
| Absorbance Max: |
278 nm |
Extinction Coefficient
(At Absorbance max): |
10200 |
| Scientific Details |
| MW: |
15 |
| Mod Code: |
/i6diPr/ |
| Absorbance Max: |
278 nm |
Extinction Coefficient
(At Absorbance max): |
10200 |
| Scientific Details |
| MW: |
15 |
| Mod Code: |
/3AmdA/ |
| Absorbance Max: |
278 nm |
Extinction Coefficient
(At Absorbance max): |
10200 |
Structure
5-Bromo-deoxyuridine is a photoreactive halogenated base that can be incorporated into oligonucleotides to crosslink them to DNA, RNA or proteins with exposure to UV light. Crosslinking is maximally efficient with light at 308 nm.
| Scientific Details |
| MW: |
78.9 |
| Mod Code: |
/55Br-dU/ |
| Absorbance Max: |
278 nm |
Extinction Coefficient
(At Absorbance max): |
9700 |
| Scientific Details |
| MW: |
78.9 |
| Mod Code: |
/i5Br-dU/ |
| Absorbance Max: |
278 nm |
Extinction Coefficient
(At Absorbance max): |
9700 |
Structure
DeoxyUridine (dU) can be substituted for dT in DNA oligonucleotides. The base can be removed by the enzyme uracil-N-deglycosylase (UNG) which renders the oligo susceptible to strand scission. One common use of this strategy is to eliminate amplified DNA and prevent cross-contamination.
| Scientific Details |
| Mod Code: |
/5deoxyU/ |
| Scientific Details |
| Mod Code: |
/ideoxyU/ |
| Scientific Details |
| Mod Code: |
/3deoxyU/ |
Structure
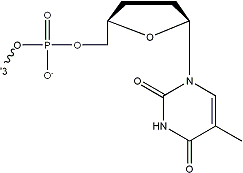
Inverted dT can be incorporated at the 3ЁЏ-end of an oligo, leading to a 3ЁЏ-3ЁЏ linkage which inhibits both degradation by 3ЁЏ exonucleases and extension by DNA polymerases.
| Scientific Details |
| Mod Code: |
/3InvdT/ |
Structure
Placing Inverted Dideoxy-T at the 5ЁЏ end of a sequence will prevent unwanted 5ЁЏ ligations.
| Scientific Details |
| Mod Code: |
/5InvddT/ |
Structure
Dideoxycytidine (ddC) is a 3ЁЏ chain terminator that prevents 3ЁЏ extension by DNA polymerases.
| Scientific Details |
| Mod Code: |
/3ddC/ |
Structure
5-Methyl deoxyCytidine when substituted for dC will increase the Tm by as much as 0.5ЁЦC per insertion. In addition, the presence of 5-Methyl dC in CpG motifs can prevent or limit unwanted immune responses that otherwise occur if oligos are administered in vivo, which is of particular importance in antisense applications.
| Scientific Details |
| MW: |
14 |
| Mod Code: |
/5Me-dC/ |
| Absorbance Max: |
277 nm |
Extinction Coefficient
(At Absorbance max): |
9000 |
| Scientific Details |
| MW: |
14 |
| Mod Code: |
/iMe-dC/ |
| Absorbance Max: |
277 nm |
Extinction Coefficient
(At Absorbance max): |
9000 |
| Scientific Details |
| MW: |
14 |
| Mod Code: |
/3Me-dC/ |
Structure
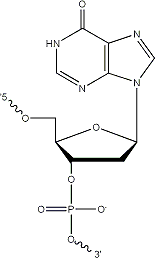
Historically, the first universal base employed was 2ЁЏ-deoxyInosine (dI). DeoxyInosine is a naturally occurring base that, while not truly universal, is less destabilizing than mismatches involving the four standard bases. Hydrogen bond interactions between dI and dA, dG, dC and dT are weak and unequal, with the result that some base-pairing bias does exist with dI:dC > dI:dA > dI:dG > dI:dT. When present in a DNA template, deoxyInosine preferentially directs incorporation of dC in the growing nascent strand by DNA polymerase.
| Scientific Details |
| Mod Code: |
/5deoxyI/ |
| Scientific Details |
| Mod Code: |
/ideoxyI/ |
| Scientific Details |
| Mod Code: |
/3deoxyI/ |
Structure
LNA bases have a modification to the ribose backbone that locks the base in the C3'-endo position, which favors RNA A-type helix duplex geometry. This modification significantly increases Tm and is also very nuclease resistant. Multiple LNA insertions can be placed in an oligo at any position except the 3'-end. Applications have been described ranging from antisense oligos to hybridization probes to SNP detection and allele specific PCR. Due to the large increase in Tm conferred by LNAs, they also can cause an increase in primer dimer formation as well as self-hairpin formation. We therefore recommend limiting the number of LNAs incorporated into a single oligo to 10 bases or less.
NOTE: TM calculations for sequences containing LNA are only estimates, as complete parameters for these calculations have yet to be published. Any applications requiring extremely accurate prediction of the TM for these probes should be reviewed with a technical support representative from Exiqon, Inc. (
http://www.exiqon.com)
Click here for more information
5-Nitroindole is currently the best universal base available. It does not favor any particular base-pairing (i.e., it does not support base-specific hydrogen bond formation), but does contribute to duplex stability through base-stacking interactions. Therefore, it is not as destabilizing to the duplex as mismatches between the standard bases. 5-Nitroindole directs random incorporation of any specific base when used as a template for DNA polymerase and partially blocks enzyme processivity.
| Scientific Details |
| MW: |
340.2 |
| Mod Code: |
/55NitInd/ |
| Extinction Coefficient: |
16000 |
| Absorbance Max: |
265 nm |
Extinction Coefficient
(At Absorbance max): |
17000 |
| Scientific Details |
| MW: |
340.2 |
| Mod Code: |
/i5NitInd/ |
| Extinction Coefficient: |
16000 |
| Absorbance Max: |
265 nm |
Extinction Coefficient
(At Absorbance max): |
17000 |
| Scientific Details |
| MW: |
340.2 |
| Mod Code: |
/35NitInd/ |
| Extinction Coefficient: |
16000 |
| Absorbance Max: |
265 nm |
Extinction Coefficient
(At Absorbance max): |
17000 |
Structure
2'-O-Methyl RNA is a naturally occurring modification of RNA found in tRNA and other small RNAs that arises as a post-transcriptional modification. Oligonucleotides can be directly synthesized that contain 2'-O-Methyl RNA. This modification increases Tm of RNA:RNA duplexes but results in only small changes in RNA:DNA stability. It is stabile with respect to attack by single-stranded ribonucleases and is typically 5 to 10-fold less susceptible to DNases than DNA. It is commonly used in antisense oligos as a means to increase stability and binding affinity to the target message.
Hydroxmethyl dC is a recently discovered modified base with a probable epigenetic role.
| Scientific Details |
| MW: |
30 |
| Mod Code: |
/55HydMe-dC/ |
| Scientific Details |
| MW: |
30 |
| Mod Code: |
/i5HydMe-dC/ |
| Scientific Details |
| MW: |
30 |
| Mod Code: |
/35HydMe-dC/ |
Structure
 IDT
IDT